2 results
Hyperendemic carbapenem-resistant Acinetobacter baumannii at a hospital in Botswana: Insights from whole-genome sequencing
- Jonathan Strysko, Tefelo Thela, Janet Thubuka, Tichaona Machiya, Jack Mkubwa, Celda Tiroyakgosi, Moses Vurayai, Kgomotso Kgomanyane, Tlhalefo Dudu Ntereke, Tshiamo Zankere, Kwana Lechiile, Teresia Gatonye, Chimwemwe Tembo, Naledi Betsi Mannathoko, Margaret Mokomane, Andries Feder, Melissa Richard-Greenblatt, David Goldfarb, Carolyn McGann, Susan Coffin, Ebbing Lautenbach, Corrado Cancedda, Dineo Bogoshi, Anthony Smith, Paul Planet
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, pp. s115-s116
-
- Article
-
- You have access Access
- Open access
- Export citation
-
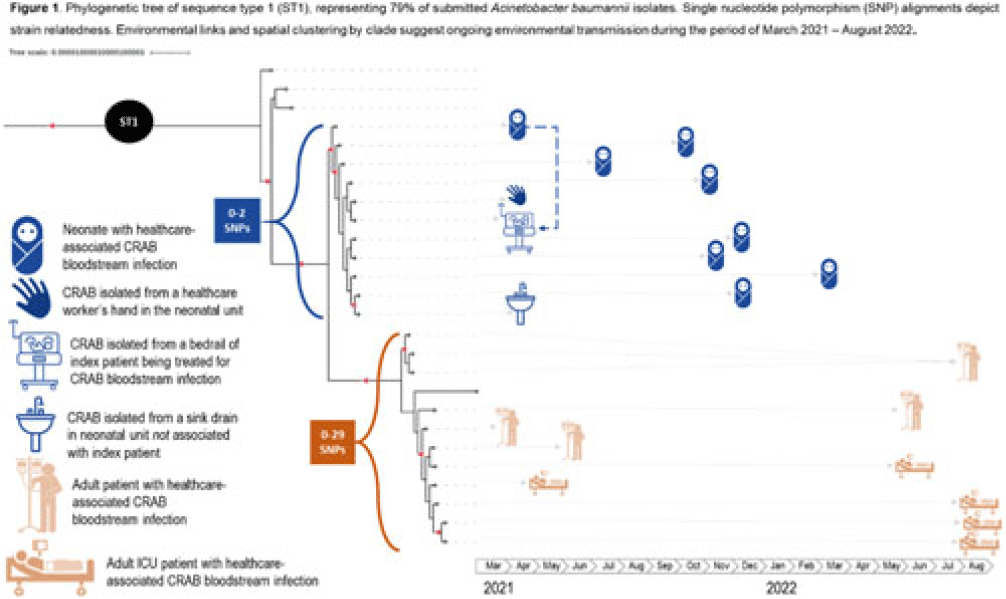
Background: Carbapenem-resistant Acinetobacter baumannii (CRAB) has emerged as a major cause of bloodstream infection among hospitalized patients in low- and middle-income countries (LMICs). CRAB infections can be difficult to treat and are devastating in neonates (~30% mortality). CRAB outbreaks are hypothesized to arise from reservoirs in the hospital environment, but outbreak investigations in LMICs seldom incorporate whole-genome sequencing (WGS). Methods: WGS (Illumina NextSeq) was performed at the National Institute for Communicable Diseases (South Africa) on 43 preserved A. baumannii isolates from a 530-bed referral hospital in Gaborone, Botswana, from March 2021–August 2022. This included 23 blood-culture isolates from 21 unique patients (aged 2 days–69 years) and 20 environmental isolates collected at the 36-bed neonatal unit in April–June 2021. Infections were considered healthcare-associated if the culture was obtained >72 hours after hospital arrival (or sooner in inborn infants). Blood cultures were incubated using an automated system (BACT/ALERT, BioMérieux) and were identified using manual methods. Environmental isolates were identified using selective or differential chromogenic media (CHROMagarTM). Taxonomic assignment, multilocus sequence typing (MLST), antimicrobial resistance gene identification, and phylogenetic analyses were performed using publicly accessible analysis pipelines. Single-nucleotide polymorphism (SNP) matrices were used to assess clonal lineage. Results: All 23 blood isolates and 5 (25%) of 20 environmental isolates were confirmed as A. baumannii; thus, 28 A. baumannii isolates were included in the phylogenetic analysis. MLST revealed that 22 (79%) of 28 isolates were sequence type 1 (ST1), including all 19 healthcare-associated blood isolates and 3 (60%) of 5 environmental isolates. Genes encoding for carbapenemases (blaNDM-1, blaOXA-23) and biocide resistance (qacE) were present in all 22 ST1 isolates; colistin resistance genes were not identified. Phylogenetic analysis of the ST1 clade demonstrated spatial clustering by hospital unit. Related isolates spanned wide ranges in time (>1 year), suggesting ongoing transmission from environmental sources (Fig. 1). An exclusively neonatal clade (0–2 SNPs) containing all 8 neonatal blood isolates was closely associated with 3 environmental isolates from the neonatal unit: a sink drain, bed rail, and a healthcare worker’s hand. Conclusions: WGS analysis of clinical and environmental A. baumannii revealed the presence of unit-specific CRAB clones, with evidence of ongoing transmission likely driven by persistent environmental reservoirs. This research highlights the potential of WGS to detect hospital outbreaks and reaffirms the importance of environmental sampling to identify and remediate reservoirs (eg, sinks) and vehicles (eg, hands and equipment) within the healthcare environment.
Disclosures: None
Colonization with extended-spectrum cephalosporin-resistant Enterobacterales (ESCrE) in hospitalized patients in Botswana
- Naledi Betsi Mannathoko, Mosepele Mosepele, Robert Gross, Rachel Smith, Ashley Styczynski, Leigh Cressman, Melissa Richard-Greenblatt, Laurel Glaser, Kevin Alby, Anne Jaskowiak, Kgotlaetsile Sewawa, Laura Cowden, Dimpho Otukile, Giacomo Paganotti, Margaret Mokomane, Warren Bilker, Ebbing Lautenbach
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 3 / Issue S2 / June 2023
- Published online by Cambridge University Press:
- 29 September 2023, p. s81
-
- Article
-
- You have access Access
- Open access
- Export citation
-
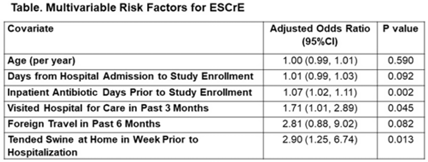
Background: The epidemiology of extended-spectrum cephalosporin-resistant Enterobacterales (ESCrE) in hospitalized patients in low- and middle-income countries (LMICs) is poorly described. Although risk factors for ESCrE clinical infection have been studied, little is known of the epidemiology of ESCrE colonization. Identifying risk factors for ESCrE colonization, which can predispose to infection, is therefore critical to inform antibiotic resistance reduction strategies. Methods: This study was conducted in 3 hospitals located in 3 districts in Botswana. In each hospital, we conducted ongoing surveillance in sequential units hospitalwide. All participants had rectal swabs collected which were inoculated onto chromogenic media followed by confirmatory testing using MALDI-TOF MS and VITEK-2. Data were collected via interview and review of the inpatient medical record on demographics, comorbidities, antibiotic use, healthcare exposures, invasive procedures, travel, animal contact, and food consumption. Participants with ESCrE colonization (cases) were compared to noncolonized participants (controls) using bivariable and multivariable analyses to identify risk factors for ESCrE colonization. Results: Enrollment occurred from January 15, 2020, to September 4, 2020, and 469 participants were enrolled. The median age was 42 years (IQR, 31–58) and 320 (68.2%) were female. The median time from hospital admission to date of sampling was 5 days (IQR, 3–12). There were 179 cases and 290 controls (ie, 38.2% of participants were ESCrE colonized). Independent risk factors for ESCrE colonization were a greater number of days on antibiotic, recent healthcare exposure, and tending swine prior to hospitalization. (Table). Conclusions: ESCrE colonization among hospitalized patients was common and was associated with several exposures. Our results suggest prior healthcare exposure may be important in driving ESCrE. The strong link to recent antibiotic use highlights the potential role of antibiotic stewardship interventions for prevention. The association with tending swine suggests that animal husbandry practices may play a role in community exposures, resulting in colonization detected at the time of hospital admission. These findings will help to inform future studies assessing strategies to curb further emergence of hospital ESCrE in LMICs.
Disclosures: None





